
Flow Cytometry (FCM) is a rapid, accurate, objective technology, whose object was fluorescent particles. The instrument can be used to detect a number of properties of a single particle (usually a cell), such as physical and biological characteristics , as well as sort out specific groups and quantified. Quantitative cellular quantification includes cell relative quantification (%), cell absolute quantification (cell/uL), cell membrane protein expression quantification (protein molecule/cells), and soluble protein absolute quantification (g/mL). The main features of flow cytometry are fast determination of cells, large sample counts, detection of rare cells, multi-parameter analysis and automation. The main use of flow cytometry
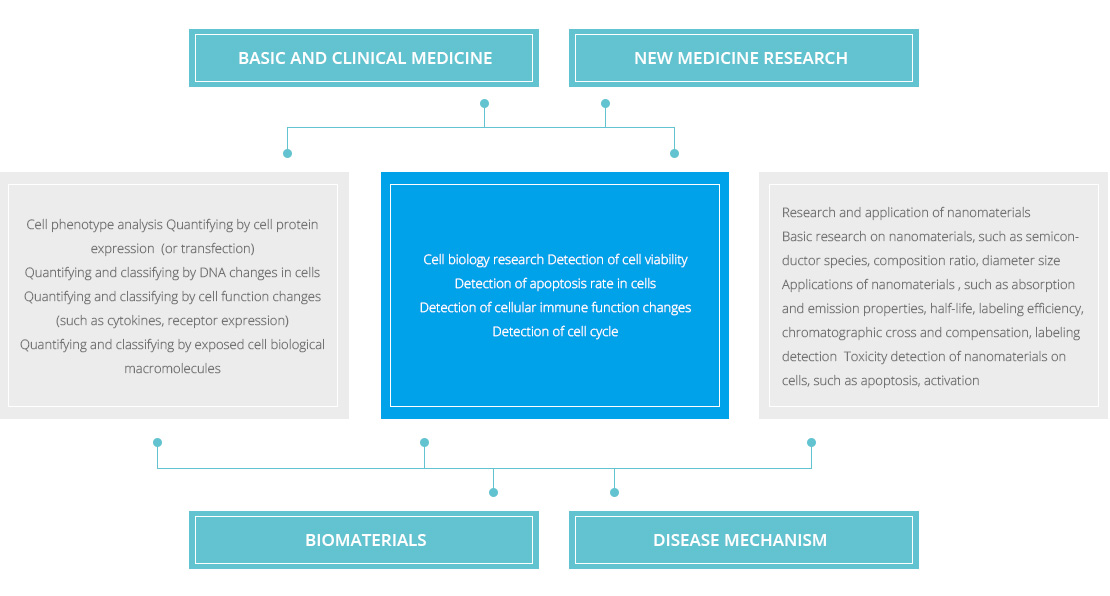
Immune status monitoring: lymphocyte subsets ratio, counting and activation of T cells, B cells, NK cells and subgroups.
Leukemia lymphoma immunophenotyping.
CD34 hematopoietic stem cell count.
Allergic reactions: basophil activation test.
Identification of autoimmune diseases: PNH, ankylosing spondylitis.
Platelet activation and functional testing: a coagulation, thrombosis tendencies.
Tumor proliferation cycle and aneuploid detection: pleural effusion, exfoliated cells.
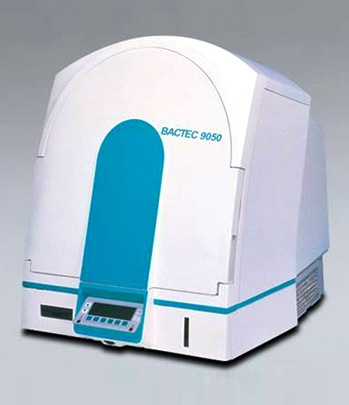
Blood cultural examination is a microbiological examination method for testing the presence of bacteria in blood samples, which is the key to effective clinical treatment. It is very important for rapid detection of sepsis, bacteremia and whether there is bacterial growth that seriously endangers the patient's life.
The detection principle of automatic blood culture instrument:
There are three detection principles of automatic blood culture instrument, namely, carbon dioxide receptors, fluorescence detection and radioactive labeling substances detection. The last method has been less used for environmental and safety reasons. The principle of the USA BD company BACTEC blood culture instrument fluorescence enhancement detection technology is the carbon dioxide released by bacteria in the metabolic process, which reacts with the sensor containing the fluorescent dye at the bottom of the flask. And then the system automatically measured the fluorescence level every 10mins, 24hrs continuously. Finally, the computer system obtains the data, along with sound, light signal alarm.
Advantages and application of automatic blood culture instrument
Culture flask of automatic blood culture instrument is the important core technology. The purpose of setting different culture bottles is mainly for the microbes on the nutritional and gas environment requirements disparity, the patient's age and physical differences and whether the use of antibiotics before cultivation. It is not only to provide different bacteria necessaries for proliferation of bacteria broth, but also contains the appropriate gas composition, to get the maximum detection of all positive specimens to prevent false negative. At present, commonly used types of flasks are standard aerobic culture bottles, standard anaerobic culture bottles, resin or activated carbon aerobic culture bottles, resin anaerobic culture bottles, resin children's culture bottles. Therefore, detection speed, accuracy and sensitivity are prominent features of the blood culture instrument.
The vacuum negative pressure extraction technology is adopted in the culture bottle, which can reduce the cost and the chance of contamination. The blood flow is uniform in blood collection to protect the patient from suffering; Blood volume collection accuracy: to avoid excessive or too little blood volume that can affect the accuracy of the test results; The closed operation of blood collection enhances the safety of medical staff. Culture bottle valid period is for 1 year, storage temperature range is 2 ℃ ~ 30 ℃; The oxygen culture bottle has been pre-filled with the necessary gas, no ventilation operation is needed, the safety of the inspectors has been improved, the cost is reduced, and the pollution is reduced also. Culture flask adopts double bar code technology, tear off one bar code to paste on the patient's report. For the query results, patients need to use a computer on the bar code reader to scan the barcode on the report which can direct access to the results of the patient and the growth curve. Usually, blood culture instrument can not only test the blood specimen, sterile fluid bacterial culture can also be used in all clinical tests, such as pleural effusion, ascites, cerebrospinal fluid, bone marrow, joint fluid, drain, bladder puncture liquid, pericardial effusion, etc.


With the completion of the human genome project, people have entered the post-genome era which is the functional genome era. The traditional sequencing methods have been unable to meet the needs of massively parallel genome sequencing, such the deep sequencing and the repeating sequencing, So that a new generation of DNA sequencing technology emerged. The new generation of sequencing technology, also known as the second generation sequencing technology includs Roche/454's sequencing technology, Illumina's Solexa as well as ABI's SOLiD sequencing technology and so on. The most significant feature of the second generation sequencing technology is the high-throughput, which can sequence hundreds of thousands to millions of DNA molecules at a time, making it easier to the transcriptome sequencing or the genome sequencing of a species. High throughput sequencing technology is that adds a joint on both sides of genomic DNA which fragmented, and then use different methods to produce millions of fixed PCR clone arrays. Each clone consists of multiple copies of a single library fragment. Next, the hybridization to primer, the enzyme mediates extension. Since all clones are on the same plane, these reactions can be carried out in massively parallel, and each of the extensions can be imaged detection by fluorescence labeled, simultaneously, thus obtaining sequencing data. DNA sequence extension and imaging detection are repeated consistently. Finally, complete sequence information of DNA can be obtained through the computer analysis. Application of High-Throughput Sequencing: We have Illumina's Miseq and Berry Genomics Corporation's NEXTseqCN500 high-throughput sequencing platforms, which can do the whole and exon genome sequencing, targeted region sequencing and so on. High-throughput sequencing technology will make genetic disease diagnosis and treatment becoming more simple and fast, and can guide individual medical care and health care on the genomic level. At the same time, the researches of biology can rely on the development of the sequencing technology. Scientists from different field can use genome sequencing of familiar species.

Sanger sequencing (first-generation sequencing) uses a DNA polymerase to extend the primers bound to the template to be determined. Until a chain terminating nucleotide is incorporated. Each sequence consisted of a set of four separate reactions, each containing all four deoxynucleotides (dNTPs) and mixed with a limited amount of a different dideoxynucleotide triphosphate (ddNTP). Since ddNTP lacks the desired 3-OH group, the extended oligonucleotide is selectively terminated at G, A, T, or C. The termination point is determined by the corresponding dideoxy in the reaction. The relative concentrations of each of the dNTPs and ddNTPs can be adjusted so that the reaction yields a set of chain-terminated products ranging from a few hundred to several thousand bases. They have a common starting point, but terminate on different nucleotides. Different fragments of different sizes can be separated by high resolution denaturing gel electrophoresis. The gel can be treated with X-ray film autoradiography or non-isotopic labeling Detection. Characteristics and Application of First-generation Sequencing:
First-generation sequencing has a high read length and a high accuracy of one-time compliance, which can handle repeat and multiply sequences well, but because of its low flux, it is only suitable for the identification of genetic disease of small sample samples. Therefore, the exon of the gene to be amplified and the specific location of the site should be clearly located, and 150 ~ 200 bp exon fragment primers upstream and downstream including this point should be designed. In addition, despite the presence of NGS, Sanger sequencing has been very economical and efficient for the detection of pathogenic genes with a clear and limited number of single-gene genetic diseases with pathogenic gene loci. So far, Sanger sequencing is still used as the golden standard for gene detection, and it is also the main method for the verification of NGS gene in the family and normal control group.
It is worth noting that the Sanger sequencing is intended to look for specific gene mutations associated with the disease. Screening of large sample cases without a clear number of candidate genes or candidate genes is difficult to accomplish, and such sequencing studies rely on NGS with high throughput sequencing capabilities. Although the Sanger sequencing has a high degree of analytical accuracy, its accuracy also depends on the sequencing instrument and the sequencing conditions. In addition, Sanger sequencing can not detect the type of gene mutation such as large fragment deletion or copy number variation, so genetic diagnosis can not be made for some related genetic diseases.

The droplet digital PCR system is subjected to microtiterization of the sample prior to conventional PCR amplification, dividing the reaction system containing the nucleic acid molecule into tens of thousands of nanostatted droplets, where each droplet is either free of a nucleic acid target Molecules, or contain one to several nucleic acid target molecules to be tested. After PCR amplification, the fluorescence signals of each droplet were analyzed one by one, the droplets of the fluorescent signal were read as 1, and the droplet without fluorescence signal was interpreted as 0, according to the Poisson distribution principle and the number of positive droplets And the ratio of the target molecule can be the starting copy number or concentration. Advantages and Applications of Droplet Digital PCR System: By diluting the sample into many parts and then counting it when a reaction occurs. The sensitivity was as high as 0.001%, significantly higher than that of the amplification block mutant system (0.1%) and Sanger sequencing (10%). While overcoming the adverse effects of nucleic acid degradation, It is very suitable for some rare samples of nucleic acid accurate detection. The absolute quantification of the true mutation rate can be calculated and the mutation rate of the target can be obtained by statistical analysis.
Droplet digital PCR is by far the most sensitive means of detecting mutations in tumor cells, which can be effectively applied to early screening of tumors. Droplet digital PCR can detect the peripheral blood samples of patients with peripheral blood circulation of tumor tissue nucleic acid analysis, compared to the traditional PCR detection method, because of a higher sensitivity, It can be used to better qualify for rare mutation detection work, from the experimental data, It was confirmed that microtiter digital PCR could achieve 0.001% mutation frequency screening. One of the main advantages of microtiter digital PCR is that the required sample size is very low, So it plays an important role in the detection of mutant / rare mutation detection. It is the best choice for the detection of drug molecules in clinical tumor samples. cfDNA real-time monitoring of tumor burden and drug resistance mutations in peripheral blood surveillance.